
Finally, through huge effort the whole team made, PmiREN was online and the team released its version 1.0 on May 6th 2019. Cheers!
In the current version, PmiREN contains 20338 miRNA loci (MIRs) belonging to 5757 families, 1365 clusters, 1668 syntenic blocks and 141327 predicted miRNA-target pairs in 88 species phylogenetically ranging from chlorophytes to angiosperms. In addition, 1587 deeply sequenced small RNA libraries were used in quantification of miRNA expression patterns, and 116 PARE-Seq libraries were employed to validate predicted miRNA-target pairs.
We made a simple comparison between PmiREN and miRBase, the most popular miRNA database. Figure A indicates the species PmiREN and miRBase harbor, respectively, where miRBase has 86 species, 28 of them both genome references and sRNA-Seq datasets are not available, while 14 of them no sRNA-Seq datasets were supported (category I and II, quite limit miRNA annotation information in miRBase). Beside, PmiREN added miRNA annotation in 44 species.
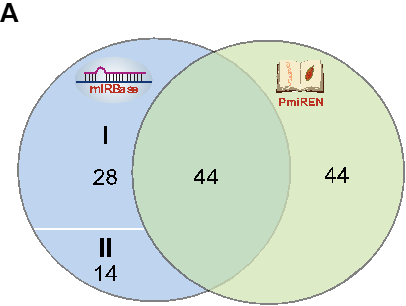
Figure B shows the comparison of miRNA candidates storing in miRBase and PmiREN. By the uniform annotation pipeline, PmiREN inherited 3722 entries (category I) from miRBase directly, changed 192 (category II) entries’ ID because of incorrect annotation, and revised 52 (category III) entries’ information such as position of mature miRNAs due to the inaccuracy annotation. Importantly, PmiREN annotated 16422 novel miRNA candidates with high confidence.
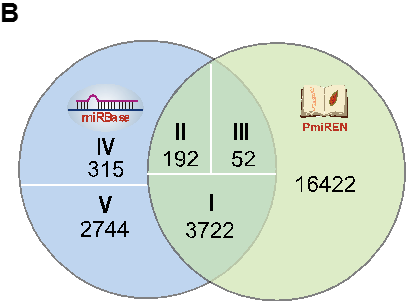
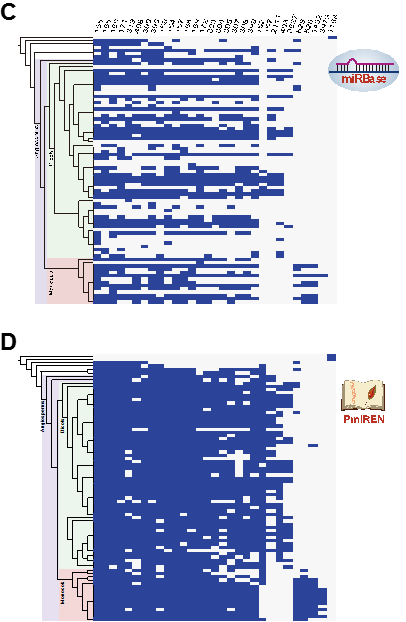
Figure C&D demonstrate the comparison of the integrity of miRNA annotation between miRBase and PmiREN. 30 most conserved miRNA families are included. Clearly, the integrity of PmiREN was greatly improved.

As of June 16th 2019 (after released 40 days), PmiREN has been visited over 30,000 times from different countries.